Croote, D., Quake, S.R. Food allergen detection by mass spectrometry: the role of systems biology. npj Syst Biol Appl. 2016 Sep 29; 2:16022.
Food allergies - a quick overview
- Up to 5% of adults and 8% of children have food allergies. [1]
- There is currently no cure for food allergies, but desensitization has been successful with oral immunotherapy.
- The most common food allergies are the "big 9": peanut, milk, egg, soy, wheat, fish, shellfish, tree nuts, and sesame.
- Protein is the allergenic component of food.
- Individuals allergic to the same food are often allergic to many of the same proteins.
- Allergenic proteins can be detected in food using analytical techniques such as enzyme-linked immunosorbent assays (ELISAs) and mass spectrometry.
[1] Sicherer, Scott H. et al. Food allergy: Epidemiology, pathogenesis, diagnosis, and treatment. J Allergy Clin Immunol, 133 (2), 291 - 307.e5. 10.1016/j.jaci.2013.11.020
Allergen detection using targeted mass spectrometry (MS)
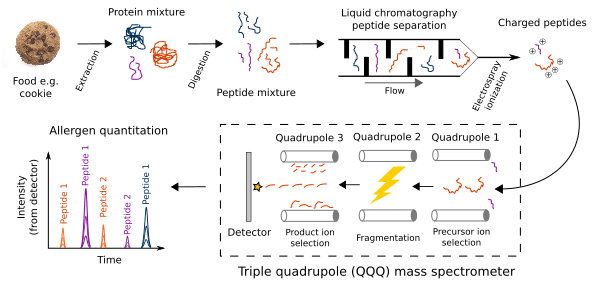
- Protein is extracted from the food
- (Not shown) Proteins are reduced and often alkylated
- The protein mixture is digested into peptides with the endopeptidase trypsin
- Peptides are temporally separated during liquid chromatography (LC) based on a peptide's relative affinity for the column's solid phase compared to the mobile phase solvent
- Peptides are ionized through electrospray ionization
- A charged peptide (known as precursor ion) is selected in the first quadrupole of the triple quadrupole (QQQ) mass spectrometer
- The precursor ion is fragmented through collision-induced dissociation into product ions in the second quadrupole
- A product ion is selected in the third quadrupole and counted by the detector
- Repeat Step 8 for other product ions
- Repeat Steps 6-9 for other precursor ions
- Calculate allergen content based on ion peak areas after adjusting for protein content, recovery, and any dilutions during sample preparation
Ready to start building a targeted method? Start by browsing allergens
About the Allergen Peptide Browser
Overview
The Allergen Peptide Browser is an interactive web tool for accelerated targeted method development utilizing published literature data along with in silico predictions. Peptide selection begins with the selection of food allergen e.g. peanut and protein e.g. Ara h 1, aided by interactive graphics conveying abundance information. Subsequently, peptides can be selected based on empirical data, in silico detection probability estimates, and/or physiochemical properties. Interactive rose plots highlight proteotypic peptides within the context of all tryptic peptides from a given protein, while physiochemical property filtering can be used to narrow the list of candidate peptides in the cases where little empirical data exists. Species specificity for each peptide is then presented based on BLAST query results of the NCBI non-redundant protein database, with warnings for instances where a peptide is not unique among food allergens. Finally, peptides can be added to a cart and exported in a comma separated value format amenable to importing into Skyline, an open-source MS software package designed to export a targeted method to a mass analyzer in native instrument format.
Peptide specificity
BLAST 2.2.31+ was used to to perform the peptide search. Only hits with 100% identity and no gaps were retained for a given peptide. Peptides associated with the same species but referencing a unique accession were grouped for easy visualization.
Filtering: The following criteria were used to exclude hits:
- Bacterial superkingdom
- Absent species name or kingdom
- Species / common name entry containing: 'synthetic construct' or 'vector'
- Title containing 'partial'
BLAST DB: Sep 20, 2016
Acknowledgements
We would like to thank the Vincent Coates Foundation Mass Spectrometry Laboratory at Stanford University for constructive discussions. This work was supported by a grant from the Simons Foundation (SFLIFE #288992 to Stephen R. Quake).
Statistics
| Total publications | 75 |
| Unique peptides detected | 822 |
| Allergenic proteins with MS data | 65 |
Literature inclusion criteria
The database has been structured around proteins recognized by the International Union of Immunological Societies (IUIS) as food allergens and belonging to one of the “big 9” food allergens, namely peanut, milk, egg, soy, wheat, tree nuts, fish, shellfish, and sesame. Empirical peptide counts were generated from the aggregation of prior publications reporting allergenic peptides in MS experiments. Inclusion has been limited to studies performing in-solution digestion with trypsin, followed with electrospray ionization (ESI) MS due to their applicability to a standardized, multiplexed quantitative workflow using selected reaction monitoring (SRM). All types of mass analyzers have been included, which has the benefits of greater confidence for truly proteotypic peptides. A full list of the references is included under the reference section.
Exporting peptides
We facilitate the development of targeted mass spectrometry methods for allergen detection by providing a cart function for saving and exporting proteotypic peptides. Transition selection and optimization along with SRM data analysis can then proceed using Skyline software [2] developed for this purpose
[2] MacLean, B. et al., Skyline: an open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics (2010) 26 (7): 966-968 10.1093/bioinformatics/btq054
Contact us
Professor Stephen R. Quake
Professor of Bioengineering, Stanford University
Head of Science at the Chan Zuckerberg Initiative
For more information, visit the group's website
Derek Croote, PhD
I am passionate about developing new technologies and therapeutics. Here are a few other places you can find me, other projects I've built, and what I'm working on:
- IgGenix - developing antibody therapeutics for allergies and other atopic conditions
- Trial Inference- a platform for accelerating clinical decisionmaking by analyzing, visualizing, and monitoring the clinical landscape
- Google Scholar